environment
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Scale
Resolution
-
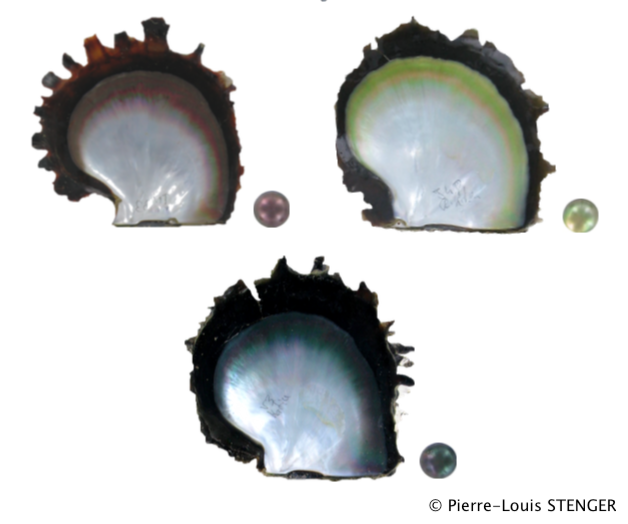

Whole genome pooled sequencing of individuals from 4 populations and 3 different color phenotype in order to uncover the genetic variants linked to color expression in the pearl oyster P. margaritifera.
-
Ces travaux ont été réalisés dans le cadre de la Directive Territoriale d'Aménagement et de Développement Durable (DTADD) portée par la Préfecture de la région ex-Aquitaine. La partie I de ces travaux porte sur les valeurs du massif forestier des Landes de Gascogne. Le massif est dépositaire d’importantes valeurs et fonctions non marchandes d’intérêt général notamment : paysagères, naturalistes, hydrologiques et climatiques. Ce rapport explique également que les modes de valorisation du territoire, autres que ceux liés à la production de bois d’œuvre et d’industrie, interfèrent étroitement avec la présence même de la forêt de production : l'activité touristique, l'arrivée de nouveaux habitants et l'économie induite, ainsi que le foncier forestier.
-
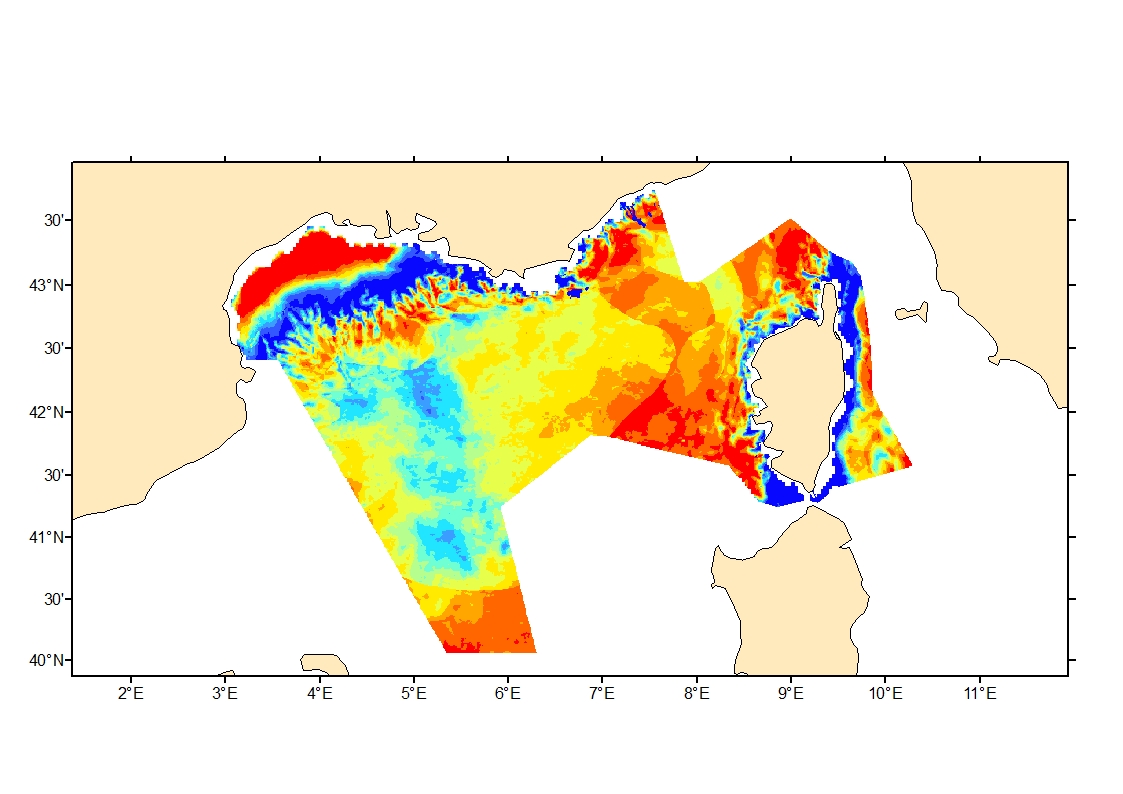
Process-driven seafloor habitat sensitivity (PDS) has been defined from the method developed by Kostylev and Hannah (2007), which takes into account physical disturbances and food availability as structuring factors for benthic communities. It is a conceptual model, relating species’ life history traits to environmental properties. Physical environment maps have been converted into a map of benthic habitat types, each supporting species communities with specific sensitivity to human pressures. It is based on two axes of selected environmental forces. The "Disturbance" (Dist) axis reflects the magnitude of change (destruction) of habitats (i.e. the stability through time of habitats), only due to natural processes influencing the seabed and which are responsible for the selection of life history traits. The "Scope for Growth" (SfG) axis takes into account environmental stresses inducing a physiological cost to organisms and limiting their growth and reproduction potential. This axis estimates the remaining energy available for growth and reproduction of a species (the energy spent on adapting itself to the environment being already taken into account). It can be related to the metabolic theory of the ecology. The process-driven sensitivity (PDS) can be seen as a risk map that combines the two previous axes and reflects the main ecological characteristics of the benthic habitats regarding natural processes. Areas with low disturbance are areas with a naturally low reworking of the sediment, allowing the establishment of a rich sessile epifauna community, with K-strategy species. Areas with low SfG means that the environmental factors, even though there are not limiting, are in lower values, i.e. that it imposes a cost for species to live. In areas combining low disturbance and low SfG, big suspension-feeder species with long life and slow growth can often be found: these species are more vulnerable in case of added disturbance.
-
L’objectif de cette étude est d’illustrer à l’aide d’indicateurs les conséquences de choix de gestion imposés par cinq scénarios socioéconomiques prospectifs appliqués à une large zone forestière pour les 60 prochaines années. Le cas d’étude choisi est la zone centrée sur la commune de Pontenx-les-Forges dans le sud-ouest de la France et couvrant 101000 hectares. Cet article présente une description de la zone d’étude et des itinéraires sylvicoles mis en œuvre par les propriétaires forestiers selon des scénarios. À l’aide d’un simulateur pilotant deux modèles de croissance, l’évolution de la zone d’étude à l’échelle de chaque parcelle est synthétisée par 9 indicateurs sur une période de 60 ans : le volume sur pied, le carbone sur pied, le volume total exploité, la valeur commerciale sur pied, le volume de l’arbre moyen, la vulnérabilité au vent et au feu, et des indices de biodiversité. Un des principaux résultats de cette étude est de montrer l’amplitude des changements pour la production et le volume sur pied : selon les scénarios les récoltes annuelles peuvent varier de 50 % dès 2030. Par conséquent, d’autres indicateurs sont impactés comme la biodiversité, la vulnérabilité au vent ou au feu. Pourtant, l’espèce dominante est maintenue et le comportement partiellement conservateur des types de propriétaires est pris en compte. En conclusion, des améliorations pour de futures simulations sont envisagées ; dans ce but, des synergies avec la télédétection sont nécessaires pour la collecte des données d’initialisation sur de larges territoires, ce qui permettra d’améliorer la précision des résultats.
-
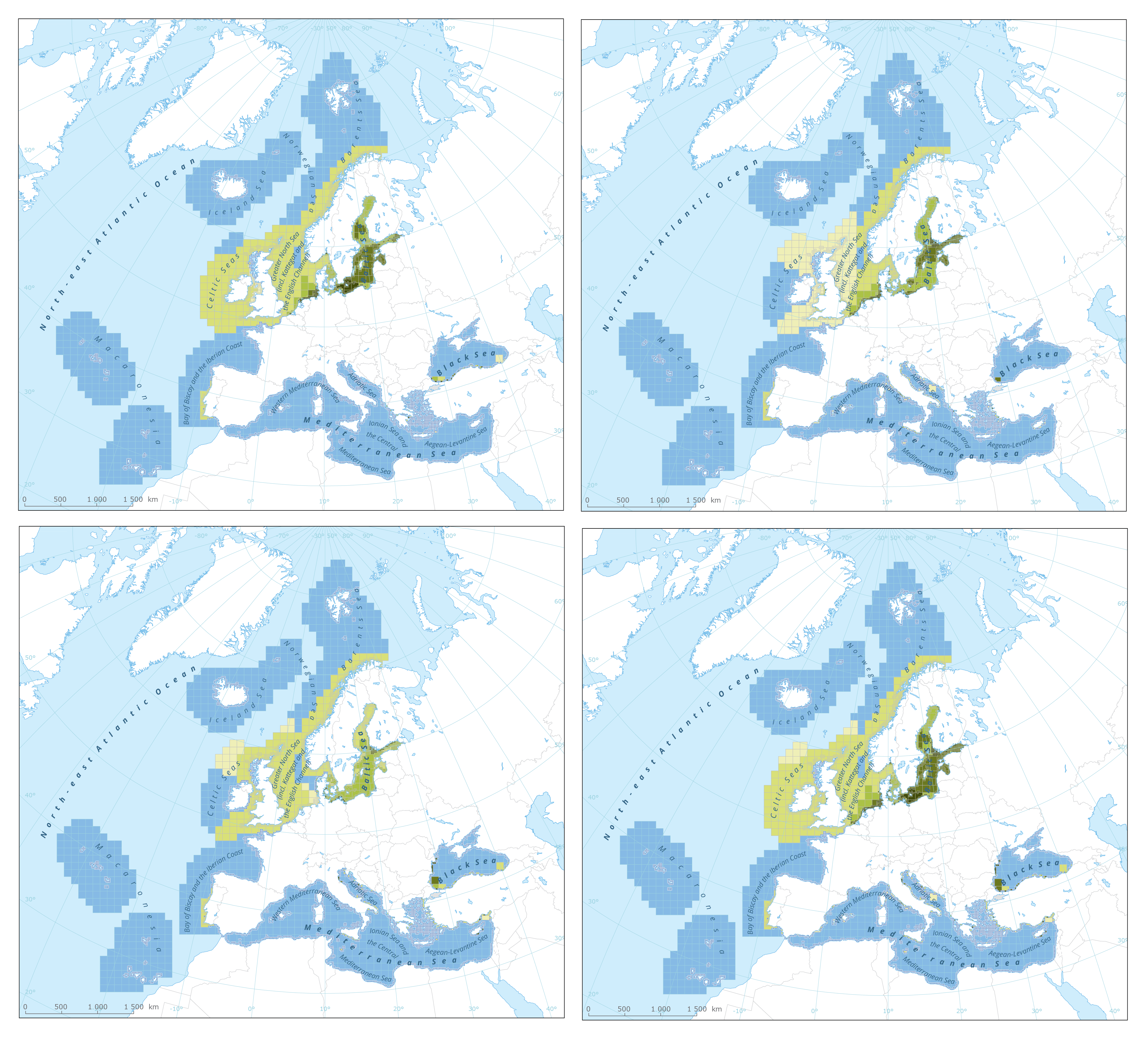
The dataset presents the results of classification of eutrophication status of the European seas using the HEAT+ tool. Eutrophication status is evaluated in five classes, where NPAhigh and NPAgood are recognised as ‘non-problem areas’ and PAmoderate, PApoor and PAbad are recognised as ‘problem areas’. Besides the overall Eutrophication status (HEAT+) are the results shown for three aspects of eutrophication: C1) Nutrient Concentrations C2) Direct Effects C3) Indirect Effects This dataset underpins the findings and cartographic representations published in the report "Nutrient enrichment and eutrophication in Europe's seas" (EEA, 2019): https://www.eea.europa.eu/publications/nutrient-enrichment-and-eutrophication-in.
-
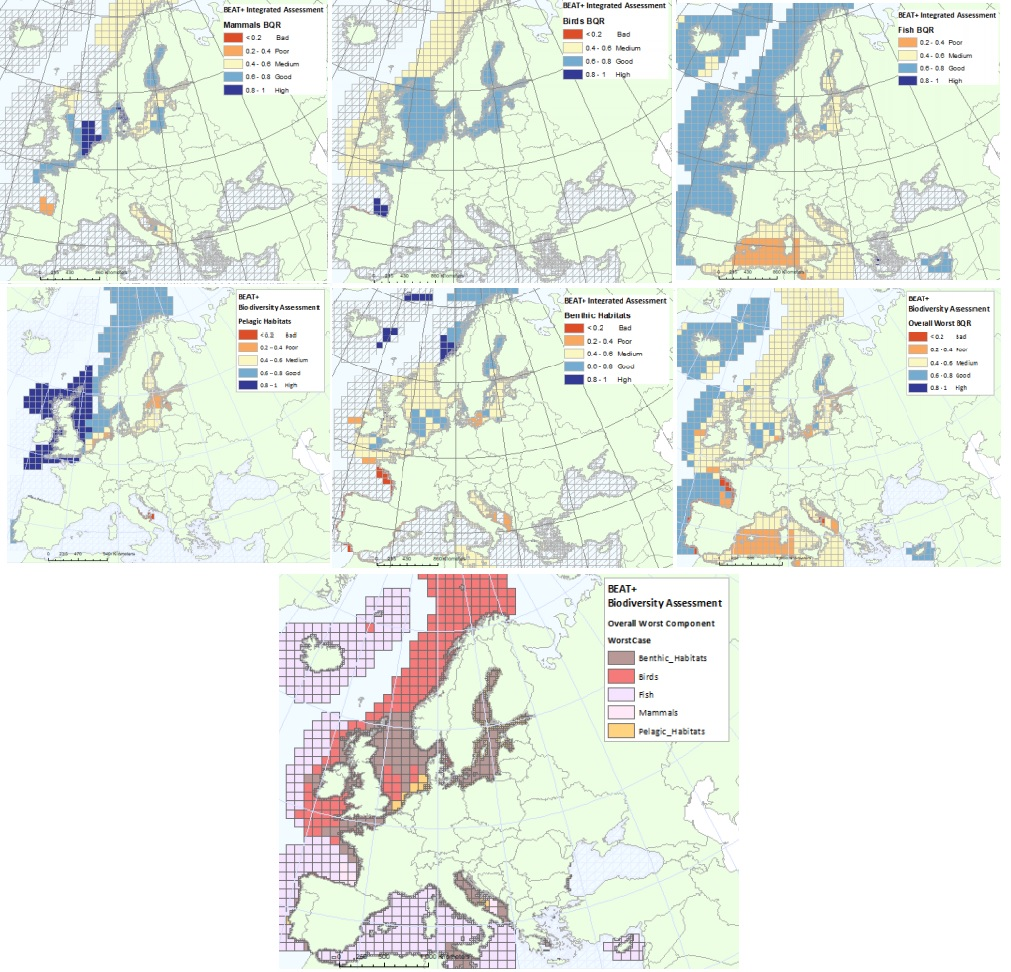
The BEAT+ tool builds on the EEA assessment tools developed and applied in the context of assessing the degree of contamination (CHASE+), eutrophication (HEAT+) and biodiversity (BEAT+) in Europe's seas. BEAT+ makes use of the same data sets and threshold values used in these assessments but recombines these in a new framework that addresses 'biodiversity condition'. BEAT+ has been designed to provide an assessment of the spatial variability of a range of biodiversity components by combining existing biodiversity indicators. The tool integrates data from normalised indicators to identify worst case status measures for different biodiversity components. The results are then linked to a standard gridE based Spatial Assessment Unit (SAU) which is used both for biodiversity and for pressures assessments (Andersen et al., 2014). These grid-based SAUs not only allow alignment of indicators for biodiversity and for pressures but provide a means for combining large assessment areas (e.g. for wide‐ranging species) with point data collected from biological surveys e.g. WFD monitoring. BEAT+ tool works by calculating a Biological Quality Ratio (BQR) which is an aggregated score of indicator outcomes within a grid square. To allow objective comparison, the indicator outcomes are normalised to a scale of 0 to 1, with five status classes at equal intervals on that scale (from Bad starting at 0, Poor at 0.2, Medium at 0.4, Good at 0.6 and High at 0.8). By this means, indicators based on different biological criteria can be aggregated in a consistent way. This metadata refers to dataset providing the results of classification of biodiversity status using the BEAT+ tool. The status is evaluated in five classes, where High and Good are recognised as ‘non-problem areas’ and Moderate, Poor and Bad are recognised as ‘problem areas’. The dataset covers: - BQR Assessment of all marine mammals combined (mainly focused on coastal and relatively stable inshore populations of seals, dolphins and porpoises) - BQR Assessment of seabirds and wading birds - BQR Assessment of commercial fish (as these have agreed targets defined on biomass and fishing mortality) - BQR Assessment of pelagic habitats - BQR Assessment of benthic habitats - BQR Assessment of worst-performing biodiversity groups - An overall synthesis of the Biological Quality Ratios (BQR) values (showing which are the worst -lowest- BQR values in each assessment grid cell. The ‘worst’ value is used here to identify the biological group most at risk, rather than averaging over all groups to avoid over-emphasis on groups with more intensive monitoring). As reference, please consult the ETC/ICM Report 3/2019: Biodiversity in Europe's seas: https://www.eionet.europa.eu/etcs/etc-icm/products/biodiversity-in-europes-seas. The indicator BEAT+ Integrated Assessment Worst Case BQR has been used in the EEA report 17/2019 "Marine Messages II": https://www.eea.europa.eu/publications/marine-messages-2.
-
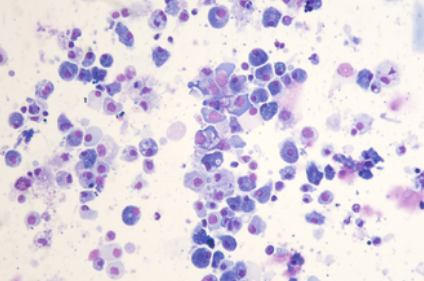
scRNA-seq reads from a Pacific oyster (Crassostrea gigas) hemocyte preparation. Hemocytes were isolated from a unique immunologically naive animal (Ifremer Standardized Animal, 18 months) and single-cell drop-seq technology was applied to 3,000 individual hemocytes.
-

Extrait de l'Atlas Aquitaine, Limousin et Poitou-Charentes sur la filière forêt-bois
-

The project was designed to explore biological rhythms in the hydrothermal vent mussel Bathymodiolus azoricus. The experiment provides the first high-resolution temporal transcriptomes of an hydrothermal species, both in situ and in the laboratory.
-

Successive infections with Vibrio harveyi were conducted in two populations of the European abalone in order to examine which genes may be involved in improved survival to the disease in the St. Malo population.
 Catalogue PIGMA
Catalogue PIGMA