Bioinformatics
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
-

Raw reads for the assembly of Gambusia holbrooki genome.
-

This dataset consists of metatranscriptomic sequencing reads corresponding to coastal micro-eukaryote communities sampled in Western Europe in 2018 and 2019.
-

Vibrio bacteria sampled from juvenile oysters and seawater collected in Thau Lagoon (Languedoc-Roussillon, France) in October 2015 during a mortality event were genotyped using hsp60, rctB, topA and mreB protein-coding genes
-
scRNA-seq reads from a Pacific oyster (Crassostrea gigas) hemocyte preparation. Hemocytes were isolated from a unique immunologically naive animal (Ifremer Standardized Animal, 18 months) and single-cell drop-seq technology was applied to 3,000 individual hemocytes.
-

DNA sequencing of Crassostrea gigas Pacific oyster spat infected in the wild with OsHV-1 virus in 4 French oyster basins (Marennes Oleron Bay, Arcachon bay, Rade de Brest and Thau lagoon).
-

WGS for Iatlantic projet ( ) for assessing past and present connectivity
-

This study aims to compare different metabarcoding sequences of commercially fished shrimps collected by tree counties on the North Brazil Shelf Large Marine Ecosystem
-

Develop parentage assignment panels using genetic fingerprinting of pearl oysters for use in commercial hatcheries and research to manage pedigrees in order to limit the risks of loss of genetic variability and increased inbreeding of commercial lines.
-

This study gathers multi-year environmental sequencing datasets generated within the French ROME pilot observatory network. It includes eDNA metabarcoding and RNA-based analyses from water samples, oyster tissues, and viral fractions collected across four French estuarine ecosystems between 2020 and 2023, supporting integrated monitoring of coastal microbiomes and microbial hazards.
-
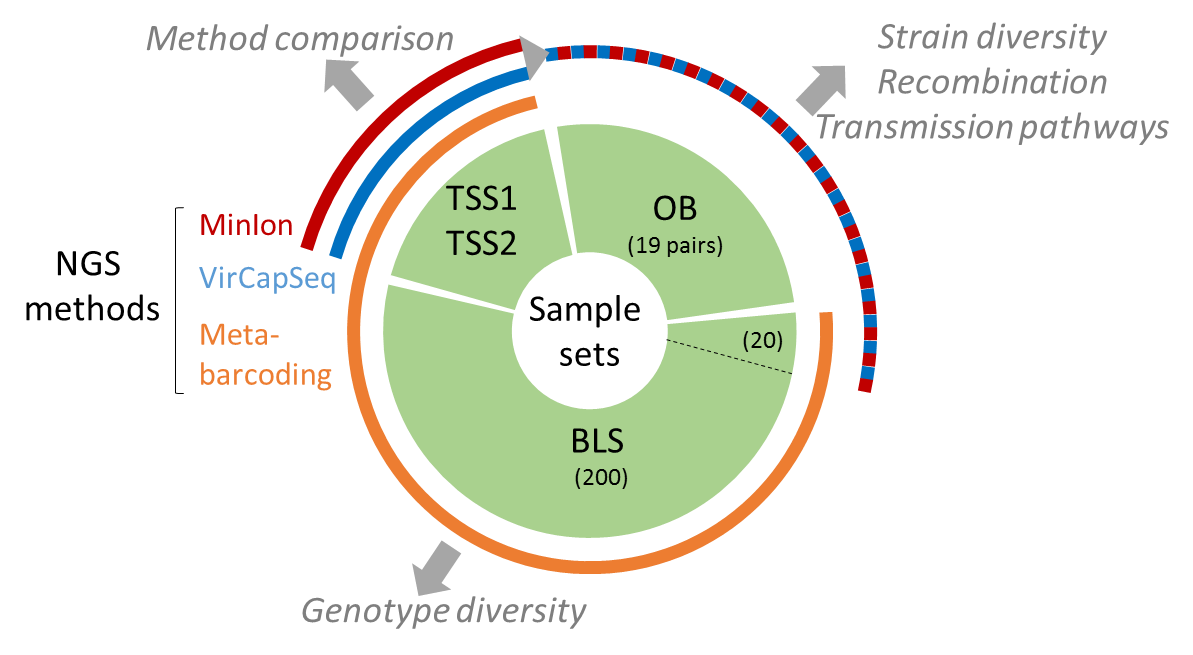
The present data set concerne metabarcoding raw reads produced using 4 different PCR targeting polymerase or capside coding region of the genoyupe I and II of norovirus. Test samples of norovirus with serial dilutions in pure water and after a bio-accumulation in oysters. Sequencing was made after VirCapSeq-VERT approach.
 Catalogue PIGMA
Catalogue PIGMA