2026
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Service types
Scale
Resolution
-

Plankton was sampled with a Continuous Underway Fish Egg Sampler (CUFES, 315µm mesh size) at 4 m below the surface, and a WP2 net (200µm mesh size) from 100m to the surface, or 5 m above the sea floor to the surface when the depth was < 100 m, in the Bay of Biscay. The full images were processed with the ZooCAM software and the embedded Matrox Imaging Library (Colas et a., 2018) which generated regions of interest (ROIs) around each individual object and a set of features measured on the object. The same objects were re-processed to compute features with the scikit-image library http://scikit-image.org. The 1, 286, 590 resulting objects were sorted by a limited number of operators, following a common taxonomic guide, into 93 taxa, using the web application EcoTaxa http://ecotaxa.obs-vlfr.fr. For the purpose of training machine learning classifiers, the images in each class were split into training, validation, and test sets, with proportions 70%, 15% and 15%. The archive contains : taxa.csv.gz Table of the classification of each object in the dataset, with columns : - objid : unique object identifier in EcoTaxa (integer number). - taxon_level1 : taxonomic name corresponding to the level 1 classification - lineage_level1 : taxonomic lineage corresponding to the level 1 classification - taxon_level2 : name of the taxon corresponding to the level 2 classification - plankton : if the object is a plankton or not (boolean) - set : class of the image corresponding to the taxon (train : training, val : validation, or test) - img_path : local path of the image corresponding to the taxon (of level 1), named according to the object id features_native.csv.gz Table of morphological features computed by ZooCAM. All features are computed on the object only, not the background. All area/length measures are in pixels. All grey levels are in encoded in 8 bits (0=black, 255=white). With columns : - area : object's surface - area_exc : object surface excluding white pixels - area_based_diameter : object's Area Based Diameter: 2 * (object_area/pi)^(1/2) - meangreyobjet : mean image grey level - modegreyobjet : modal object grey level - sigmagrey : object grey level standard deviation - mingrey : minimum object grey level - maxgrey : maximum object grey level - sumgrey : object grey level integrated density: object_mean*object_area - breadth : breadth of the object along the best fitting ellipsoid minor axis - length : breadth of the object along the best fitting ellipsoid majorr axis - elongation : elongation index: object_length/object_breadth - perim : object's perimeter - minferetdiam : minimum object's feret diameter - maxferetdiam : maximum object's feret diameter - meanferetdiam : average object's feret diameter - feretelongation : elongation index: object_maxferetdiam/object_minferetdiam - compactness : Isoperimetric quotient: the ration of the object's area to the area of a circle having the same perimeter - intercept0, intercept45 , intercept90, intercept135 : the number of times that a transition from background to foreground occurs a the angle 0ø, 45ø, 90ø and 135ø for the entire object - convexhullarea : area of the convex hull of the object - convexhullfillratio : ratio object_area/convexhullarea - convexperimeter : perimeter of the convex hull of the object - n_number_of_runs : number of horizontal strings of consecutive foreground pixels in the object - n_chained_pixels : number of chained pixels in the object - n_convex_hull_points : number of summits of the object's convex hull polygon - n_number_of_holes : number of holes (as closed white pixel area) in the object - roughness : measure of small scale variations of amplitude in the object's grey levels - rectangularity : ratio of the object's area over its best bounding rectangle's area - skewness : skewness of the object's grey level distribution - kurtosis : kurtosis of the object's grey level distribution - fractal_box : fractal dimension of the object's perimeter - hist25, hist50, hist75 : grey level value at quantile 0.25, 0.5 and 0.75 of the object's grey levels normalized cumulative histogram - valhist25, valhist50, valhist75 : sum of grey levels at quantile 0.25, 0.5 and 0.75 of the object's grey levels normalized cumulative histogram - nobj25, nobj50, nobj75 : number of objects after thresholding at the object_valhist25, object_valhist50 and object_valhist75 grey level - symetrieh :index of horizontal symmetry - symetriev : index of vertical symmetry - skelarea : area of the object skeleton - thick_r : maximum object's thickness/mean object's thickness - cdist : distance between the mass and the grey level object's centroids features_skimage.csv.gz Table of morphological features recomputed with skimage.measure.regionprops on the ROIs produced by ZooCAM. See http://scikit-image.org/docs/dev/api/skimage.measure.html#skimage.measure.regionprops for documentation. inventory.tsv Tree view of the taxonomy and number of images in each taxon, displayed as text. With columns : - lineage_level1 : taxonomic lineage corresponding to the level 1 classification - taxon_level1 : name of the taxon corresponding to the level 1 classification - n : number of objects in each taxon group map.png Map of the sampling locations, to give an idea of the diversity sampled in this dataset. imgs Directory containing images of each object, named according to the object id objid and sorted in subdirectories according to their taxon.
-

The Mediterranean Sea is a natural laboratory to address questions about the formation and evolution of continental margins and the relationship between surface and deep processes. Different regional to local events have influenced the Neogene stratigraphic evolution of the Valencia and Menorca basins. The evaporites deposited during the Messinian Salinity Crisis (MSC) strongly impact its sedimentary and geomorphological evolution. Here we present a compilation of the main regional seismic stratigraphic markers from the continental platform to the deep sea. We provide in xyz format (z in second twt) the original picking files, (not interpolated) and interpolated grid of: i) the top of the Mesozoic formation, the base of the Neogene formations including the early Miocene volcanic features, ii) the top Burdigalian, Langhian, and Serravallian seismic horizons, iii) the seismic horizons related to the Messinian Salinity Crisis, iv) the Pliocene and Pleistocene seismic horizons v) the depth of the Seafloor. The available reflection seismic dataset results from the compilation and processing of vintage seismic profiles of previous works and from the Instituto Geologico y Minero de Espana (IGME). This compilation is currently the first available in literature and provides a useful contribution to the scientific community working on sedimentary, tectonics and geodynamics within the Western Mediterranean basins.
-

The network was initiated by IFREMER from 1993 to 2009 (under the acronym REMORA) to study the rearing performance of the Pacific oyster Crassostrea gigas at a national scale. To do so, the network monitored annually the mortality and growth of standardized batches of 18-month-old oysters. Starting in 1995, the monitoring of the rearing performance of 6-month-old oyster spat was integrated into this network. These sentinel batches were distributed simultaneously each year on 43 sites and were monitored quarterly. These sites were distributed over the main French oyster farming areas and allowed a national coverage of the multiannual evolution of oyster farming performances. Most of the sites were located on the foreshore at comparable levels of immersion. Field studies were carried out by the "Laboratoires Environnement Ressources" (LER) for the sites included in their geographical area of investigation. Following the increase in spat mortality in 2008, the network evolved in 2009 (under the acronym RESCO). From this date, the network selected 13 sites among the 43 sites previously monitored in order to increase the frequency of visits (twice a month) and the number of sentinel batches. More precisely, sentinel batches of oysters corresponding to different origins (wild or hatchery, diploid or triploid) and to two rearing age classes (spat or 18-month-old adults) were selected. The monitoring of environmental variables (temperature, salinity) associated with the 13 sites was also implemented. The actions of the network have thus contributed to disentangle the biotic and abiotic parameters involved in mortality phenomena, taking into account the different compartments (environment / host / infectious agents) likely to interact with the evolution of oyster rearing performance. Finally, since 2015, the network has merged the RESCO and VELYGER networks to adopt the acronym ECOSCOPA. The general objective of this current network is to analyze the causes of spatio-temporal variability of the main life traits (Larval stage - Recruitment - Reproduction - Growth - Survival - Cytogenetic abnormalities) of the cupped oyster in France and to follow their evolution on the long term in the context of climate change. To do this, the network proposes a regular spatio-temporal monitoring of the major proxies of the life cycle of the oyster, organized in three major thematic groups: (1) proxies related to growth, physiological tolerance and survival of experimental sentinel populations over 3 age classes: (2) proxies related to reproduction, larval phase and recruitment of the species throughout its natural range in France, and: (3) proxies related to environmental parameters essential to the species (weather conditions, temperature, salinity, pH, turbidity, chlorophyll a and phytoplankton) at daily or sub-hourly frequencies. Working in a geographical network associating several laboratories, ECOSCOPA provide these monitoring within 8 sites selected among the previous ones to ensure the continuity of the data acquisition. Today, these 8 sites are considered as ecosystems of common interest, contrasted, namely : - The Thau lagoon - The Arcachon basin - The Marennes Oléron basin - The Bourgneuf Bay - The bay of Vilaine - The bay of Brest - The bay of Mont Saint Michel - The bay of Veys The ECOSCOPA network is therefore one of the relevant monitoring tools on a national scale, allowing to objectively measure through different proxies the general state of health of cultivated and wild oyster populations, and this for the different sensitive phases of their life cycle. This network aims at allowing a better evaluation, on the long term, of the biological risks incurred by the sector but also by the ecosystems, in particular under the increasing constraint of climatic and anthropic changes. Figure : Sites monitored by the ECOSCOPA network
-

Within the ESA Coastal Blue Carbon project, the LIENSs laboratory contributed drone-derived products, SP80 ground survey points (with longitude, latitude, and plant data) and biomass measurements to test classification models for mapping salt marsh vegetation (e.g., Esnandes) and associated biomass/carbon stocks. Links to associated datasets are provided at the bottom of this sheet.
-

The ORHAGO campaigns (Observation of the benthic aquatic resources of the Golfe de Gascogne) are designed to collect data on the composition, distribution and change in relative abundance of benthic fish fauna on the continental shelf (depth <100m) in November to December on a yearly basis. The ORHAGO survey was initiated in 2007 with the objective of developing a fishery-independent abundance index for flatfish species, with a particular focus on the common sole (Solea solea) of the Bay of Biscay. In accordance with the ICES-agreed gear for flatfish abundance surveys, ORHAGO employs a 4-meter-beam trawl with a chain mat, 50-millimeter mesh in the net, and 40-millimeter mesh in the cod-end. The sampling plan was designed to ensure full coverage of the common sole habitat in the Bay of Biscay during a period (November-December) for which fish behavior and distribution was suitable for obtaining an unbiased abundance index (young fish move offshore when coastal waters become colder and before the concentrations of the spawning season). The sampling design is a systematic sampling with 49 reference stations. The sampling design was validated in 2013 by the ICES Working Group on Beam Trawl Surveys (WGBEAM). Since 2013, the ORHAGO survey has been used to assess the status of the Bay of Biscay common sole stock (WGBIE, Working Group for the Bay of Biscay and the Iberian Waters Ecoregion).
-

A total of 210 points in 2024 (28 May, 18 april, 13 September, 19 September, and 25 October) and 195 points in 2025 (8 April, 18 and 29 April, 3 June, 17 June, and 4 July) were surveyed with an SP80 DGPS as part of the ESA Coastal Blue Carbon project. Additionally, 174 and 160 points obtained from photointerpretation of multispectral drone orthomosaics from 19 September 2024 and 18 September 2025, respectively, were recorded. At each point, the vegetation type was specified.
-
Serveur wms public de l'Ifremer, Accès aux données du Sismer
-
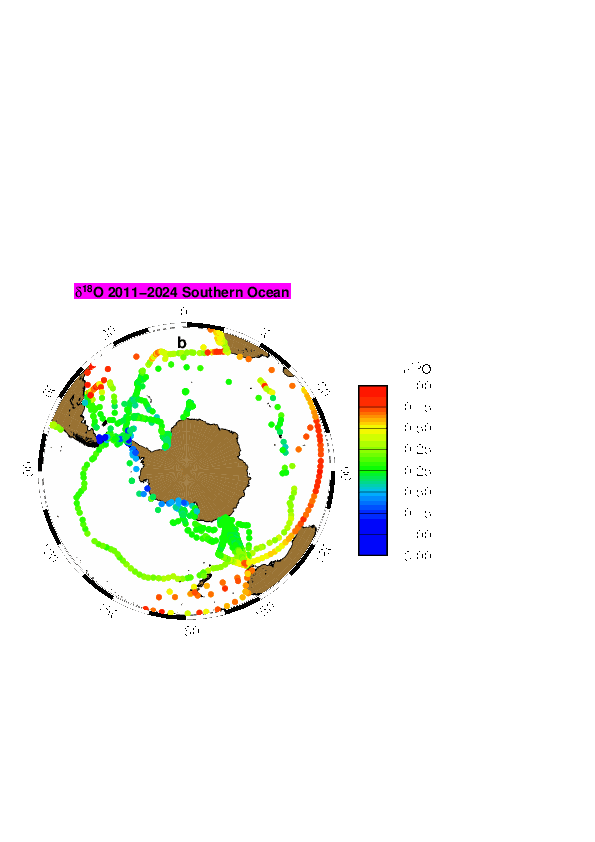
LOCEAN has been in charge of collecting sea water for the analysis of water isotopes on a series of cruises or ships of opportunity mostly in the equatorial Atlantic, in the North Atlantic, in the southern Indian Ocean, in the southern Seas, Nordic Seas, and in the Arctic. The LOCEAN data set of the oxygen and hydrogen isotope (δ18O and δD) of marine water covers the period 1998 to 2019, but the effort is ongoing. Most data prior to 2010 (only δ18O) were analyzed using isotope ratio mass spectrometry (Isoprime IRMS) coupled with a Multiprep system (dual inlet method), whereas most data since 2010 (and a few earlier data) were obtained by cavity ring down spectrometry (CRDS) on a Picarro CRDS L2130-I, or less commonly on a Picarro CRDS L2120-I. Occasionally, some data were also run by Marion Benetti on an Isoprime IRMS coupled to a GasBench (dual inlet method) at the university of Iceland (Reykjavik). On the LOCEAN Picarro CRDS, most samples were initially analyzed after distillation, but since 2016, they have often been analyzed using a wire mesh to limit the spreading of sea salt in the vaporizer. Some of the samples on the CRDS were analyzed more than once on different days, when repeatability for the same sample was not sufficient or the daily run presented a too large drift. Accuracy is best when samples are distilled, and for δD are better on the Picarro CRDS L2130-I than on the Picarro CRDS L2120-I. Usually, we found that the reproducibility of the δ18O measurements is within ± 0.05 ‰ and of the δD measurements within ± 0.30 ‰, which should be considered an upper estimate of the error on the measurement on a Picarro CRDS. The water samples were kept in darkened glass bottles (20 to 50 ml) with special caps, and were often (but not always) taped afterwards. Once brought back in Paris, the samples were often stored in a cold room (with temperature close to 4°C), in particular if they were not analyzed within the next three months. There is however the possibility that some samples have breathed during storage. We found it happening on a number of samples, more commonly when they were stored for more than 5 years before being analyzed. We also used during one cruise bottles with not well-sealed caps (M/V Nuka Arctica in April 2019), which were analyzed within 3 months, but for which close to one third of the samples had breathed. We have retained those analyses, but added a flag ‘3’ meaning probably bad, at least on d-excess (outside of regions where sea ice forms or melts, for the analyses done on the Picarro CRDS, excessive evaporation is usually found with a d-excess criterium (which tends to be too low); for the IRMS analyses, it is mostly based when excessive scatter is found in the S- δ18O scatter plots or between successive data, in which case some outliers were flagged at ‘3’). In some cases when breathing happened, we found that d-excess can be used to produce a corrected estimate of δ18O and δD (Benetti et al., 2016). When this method was used a flag ‘1’ is added, indicating ‘probably good’ data, and should be thought as not as accurate as the data with no ‘correction’, which are flagged ‘2’ or ‘0’. We have adjusted data to be on an absolute fresh-water scale based on the study of Benetti et al. (2017), and on further tests with the different wire meshes used more recently. We have also checked the consistency of the runs in time, as there could have been changes in the internal standards used. On the Isoprime IRMS, it was mostly done using different batches of ‘Eau de Paris’ (EDP), whereas on the Picarro CRDS, we used three internal standards kept in metal tanks with a slight overpressure of dry air). The internal standards have been calibrated using VSMOW and GISP, and were also sent to other laboratories to evaluate whether they had drifted since the date of creation (as individual sub-standards have typically stored for more than 5-years). These comparisons are still not fully statisfactory to evaluate possible drifts in the sub-standards. Version V5 contains only one global file (ALL-Wisotopes-V5). However, up to version V4, individual files corresponded to regional subsets : - SO: Southern Ocean including cruise station and surface data mostly from 2017 in the Weddell Sea (WAPITI Cruise JR160004, DOI:10.17882/54012), as well as in the southern Ocean south of 20°S - SI: OISO cruise station and surface data in the southern Indian Ocean (since 1998) (DOI:10.18142/228) - EA: 20°N-20°S cruise station and surface data (since 2005), in particular in the equatorial Atlantic from French PIRATA (DOI:10.18142/14) and EGEE cruises (DOI:10.18142/95) - NA: 20°N-72°N station and surface data, mostly in the North Atlantic from Oceanographic cruises as well as from ships of opportunity (this includes in particular OVIDE cruise data since 2002 (DOI:10.17882/46448), CATARINA, BOCATS1 and BOCATS2 (PID2019-104279GB-C21/AEI/10.13039/501100011033) cruises funded by the Spanish Research Agency, RREX2017 2017 cruise data (DOI:10.17600/17001400), SURATLANT data set since 2011 (DOI:10.17882/54517), Nuka Arctica and Tukuma Arctica data since 2012, STRASSE (DOI:10.17600/12040060) and MIDAS cruise data in 2012-2013, as well as surface data from various ships of opportunity since 2012) - NS: Nordic Sea data from cruises in 2002-2018 - AS: Arctic Ocean north of 72°N, in particular from two Tara cruises (in 2006-2008 and 2013) and expeditions since 2020 - PM: miscellaneous data in tropical Pacific, Indian Ocean, Mediterranean Sea and Black Sea In some regions, such as in the Indian Ocean, it is valuable to combine different subsets to have the full data distribution. The files are in csv format reported, and starting with version V1, it is reported as: - Cruise name, station id, bottle number, day, month, year, hour, minute, latitude, longitude, pressure (db), temperature (°C), it, salinity (pss-78), is, dissolved oxygen (micromol/kg), io2, δ18O, iO, d D, iD, d-excess, id, method type - Temperature is an in situ temperature - Salinity is a practical salinity it, is, io2, iO, iD, id are quality indices equal to: - 0 no quality check (but presumably good data) - 1 probably good data - 2 good data - 3 probably bad data - 4 certainly bad data - 9 missing data (and the missing data are reported with an unlikely missing value) The method type is 1 for IRMS measurements, 2 for CRDS measurement of a saline water sample, 3 for CRDS measurement of a distilled water sample.
-
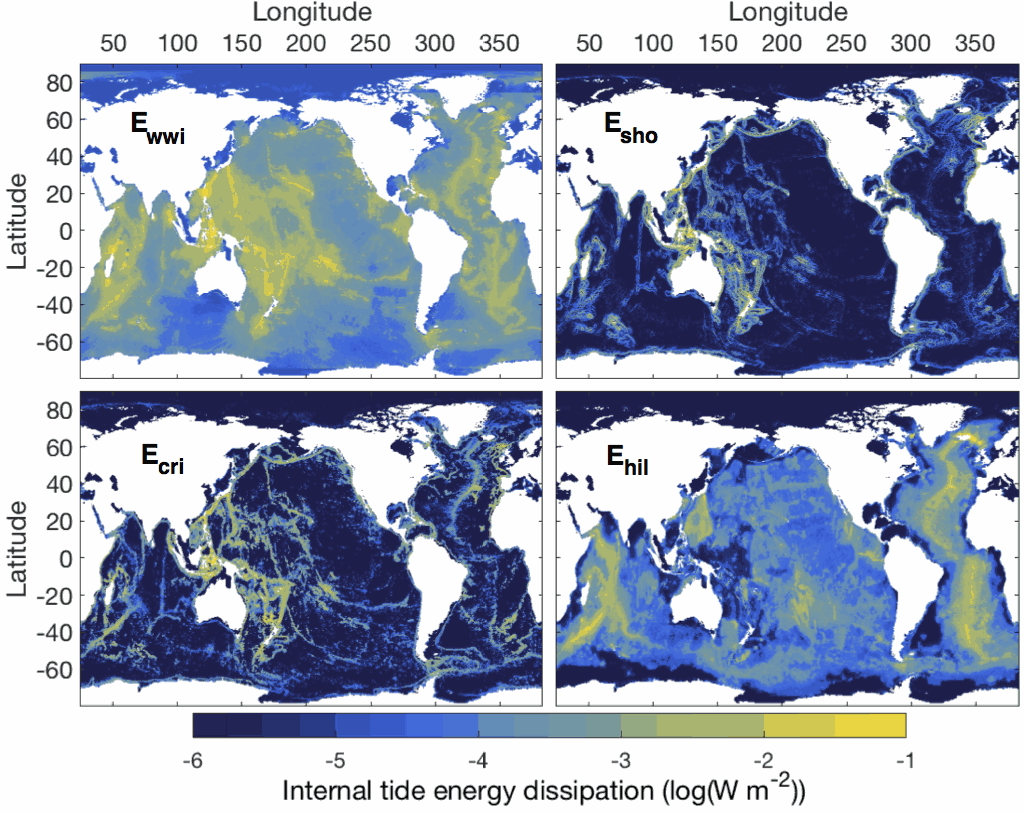
This dataset comprises two netcdf files. The first file contains the six global two-dimensional maps necessary to implement the tidal mixing parameterization presented in de Lavergne et al. (2020). Four power fields (E_wwi, E_sho, E_cri and E_hil) represent depth-integrated internal tide energy dissipation, with units of Watts per square meter. Each power field corresponds to a specific dissipative process and associated vertical structure of turbulence production. The two remaining fields, H_cri and H_bot, are decay heights (with units of meters) that enter the vertical structures of the E_cri and E_hil components, respectively. The second file contains three-dimensional fields of turbulence production (with units of Watts per kilogram) obtained by application of the parameterization to the WOCE global hydrographic climatology. The file includes the total turbulence production (epsilon_tid), its four components (epsilon_wwi, epsilon_sho, epsilon_cri, epsilon_hil), and the underlying hydrographic fields, as a function of longitude, latitude and depth. All maps have a horizontal resolution of 0.5º. Detailed documentation of the parameterization can be found in the following publication: de Lavergne, C., Vic, C., Madec, G., Roquet, F., Waterhouse, A.F., Whalen, C.B., Cuypers, Y., Bouruet-Aubertot, P., Ferron, B., Hibiya, T. A parameterization of local and remote tidal mixing. Journal of Advances in Modeling Earth Systems, 12, e2020MS002065 (2020). https://doi.org/10.1029/2020MS002065
-

Plankton was imaged with the PlanktoScope in different oceanic regions using different nets and protocol of conservation. This dataset aims to serve as reference for taxonomic identification with the PlanktoScope across 256 plankton taxa from 20µm to 300µm. Reference dataset can also serve as learning set for prediction in Ecotaxa (https://ecotaxa.obs-vlfr.fr/prj/15535). The full images were processed and segmented with the PlanktoScope around each individual. A set of associated features were measured on the objects with skimage.measure. All objects were classified into 256 different classes using the web application EcoTaxa (http://ecotaxa.obs-vlfr.fr). The following dataset corresponds to the 169, 149 objects and their calculated features. The different files provide information about the features of the objects, their taxonomic identification as well as the raw images. taxa.csv.gz Table of the classification of each object in the dataset, with columns: - object_id: unique object identifier in EcoTaxa. - annotation_category: taxonomic name corresponding to the last level of hierarchy - annotation_hierarchy: taxonomic lineage to the category - set: class of the image corresponding to the taxon - img_file_name: local path of the image corresponding to the taxon, named according to the object id features_native.csv.gz Table of morphological features computed by PlanktoScope. All features are computed on the object only, not the background. All area/length measures are in pixels. - object_id: unique object identifier in Ecotaxa And 33 features: - width: width of the smallest rectangle enclosing the object (pixel) - height: height of the smallest rectangle enclosing the object (pixel) - bx: X coordinates of the top left point of the smallest rectangle enclosing the object (pixel) - by: Y coordinates of the top left point of the smallest rectangle enclosing the object (pixel) - circ.: circularity of the object ((4∗π ∗Area)/Perim^2) [0-1] - area_exc: Surface area of the object excluding holes (pixel2) - area: Surface area of the object (pixel2) - %area: Percentage of object’s surface area that is comprised of holes - major: Length of the primary axis of the best fitting ellipse for the object (pixel) - minor: Length of the secondary axis of the best fitting ellipse for the object (pixel) - y: Y position of the center of gravity of the object (pixel) - x: X position of the center of gravity of the object (pixel) - convex_area: The area of the smallest polygon within which all points in the object fit (pixel2) - perim.: The length of the outside boundary of the object (pixel) - elongation: elongation index (major/minor) - perimareaexc: index of the relative complexity of the perimeter (perim/area_exc) - perimmajor: Index of the relative complexity of the perimeter (perim/major) - circex: Circularity of object excluding white pixels ((4 ∗ π ∗ Area_exc)/perim 2) - angle: Angle between the primary axis and a line parallel to the x-axis of the image - bounding_box_area: Area of the smallest box containing the object (pixel2) - eccentricity: Eccentricity of the ellipse that has the same second-moments as the region. Ratio of the focal distance of the ellipse over the major axis length [0-1] - equivalent_diameter: The diameter of a circle with the same area as the object (pixel) - euler_number: Euler characteristic of the set of non-zero pixels. Computed as number of connected components subtracted by number of holes - extent: Ratio of pixels in the object to pixels in the total bounding box - local_centroid_col: Horizontal coordinate of the center of mass of the object (pixel) - local_centroid_row: Vertical coordinate of the center of mass of the object (pixel) - solidity: Ratio of pixels in the object to pixels of the convex hull image (area / convex_area) - meanhue: Mean base color of the object in hue scale (0-360) - meansaturation: Mean saturation of the object [0-100] - meanvalue: Mean brightness of the object [0-100] - stdhue: Standard deviation of base color - stdsaturation: Standard deviation of saturation - stdvalue: Standard deviation of brightness inventory.tsv Tree view of the taxonomy and number of images in each taxon, displayed as text. With columns : - annotation_hierarchy: taxonomic lineage - annotation_category: name of the taxon - n: number of objects in each taxon group map.png Map of the sampling locations, to give an idea of the diversity sampled in this dataset. imgs Directory containing images of each object, named according to the object id object_id and sorted in subdirectories according to their taxon.
 Catalogue PIGMA
Catalogue PIGMA