environment
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Scale
Resolution
-

Première utilisation du sol, devant l'agriculture et loin devant l'urbanisme, la forêt couvre 45 % du territoire aquitain. La région se caractérise par la domination d'une essence, le pin maritime. Celui-ci couvre plus de la moitié de la surface forestière régionale. Outre sa valeur patrimoniale, cette forêt génère une activité économique qui représente environ 3 milliards d'euros. Ce secteur forêt-bois est donc un formidable gisement d'emplois, principalement en milieu rural. Cet espace occupé par la forêt attise néanmoins des convoitises pour différents types d'usage: l'urbanisation, les installations photovoltaïques ou encore l'agriculture.
-
L’objectif de cette étude est d’illustrer à l’aide d’indicateurs les conséquences de choix de gestion imposés par cinq scénarios socioéconomiques prospectifs appliqués à une large zone forestière pour les 60 prochaines années. Le cas d’étude choisi est la zone centrée sur la commune de Pontenx-les-Forges dans le sud-ouest de la France et couvrant 101000 hectares. Cet article présente une description de la zone d’étude et des itinéraires sylvicoles mis en œuvre par les propriétaires forestiers selon des scénarios. À l’aide d’un simulateur pilotant deux modèles de croissance, l’évolution de la zone d’étude à l’échelle de chaque parcelle est synthétisée par 9 indicateurs sur une période de 60 ans : le volume sur pied, le carbone sur pied, le volume total exploité, la valeur commerciale sur pied, le volume de l’arbre moyen, la vulnérabilité au vent et au feu, et des indices de biodiversité. Un des principaux résultats de cette étude est de montrer l’amplitude des changements pour la production et le volume sur pied : selon les scénarios les récoltes annuelles peuvent varier de 50 % dès 2030. Par conséquent, d’autres indicateurs sont impactés comme la biodiversité, la vulnérabilité au vent ou au feu. Pourtant, l’espèce dominante est maintenue et le comportement partiellement conservateur des types de propriétaires est pris en compte. En conclusion, des améliorations pour de futures simulations sont envisagées ; dans ce but, des synergies avec la télédétection sont nécessaires pour la collecte des données d’initialisation sur de larges territoires, ce qui permettra d’améliorer la précision des résultats.
-

Présentation des entreprises, cartographie du risque et consignes en cas d'alerte pour les populations des communes de la Presqu'île d'Ambès. Plaquettes 4 ou 8 pages (avec cartographie)
-
Massif forestier des Landes de Gascogne : Guide pour la prise en compte du risque incendie de forêt.

Ce guide a pour vocation d'informer sur les caractéristiques du risque incendie de forêt propres au massif des Landes de Gascogne, de définir les modalités de prise en compte du risque dans les documents d'urbanisme et de regrouper l'ensemble des réglementations et recommandations ayant trait à la protection contre les incendies. Il a été élaboré par l'Etat en partenariat avec l'Association des Maires des Landes et les organismes concernés par cette problématique.
-
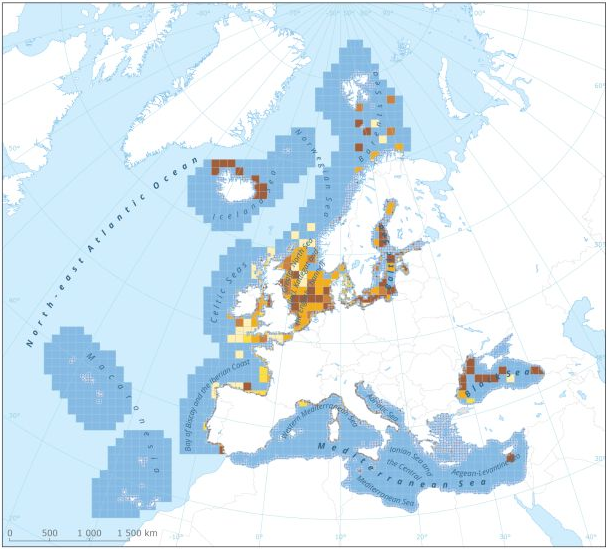
This dataset presents the resulting assessment grid (based on the EEA reference grid) with the classification of chemical status of the transitional, coastal and marine waters in the context of the Water Framework Directive (WFD) and the Marine Strategy Framework Directive (MSFD). This classification has been performed using the CHASE+ tool, with classifications of the matrices ‘water’, ‘sediment’ and ‘biota’ and indicators of ‘biological effects’, as well as an integrated classification of chemical status, combining results of all matrices. The chemical status is evaluated in five classes, where NPAhigh and NPAgood are recognised as ‘non-problem areas’ and PAmoderate, PApoor and PAbad are recognised as ‘problem areas’. This is the assessment made excluding concentrations of mercury (Hg). The overall area of interest used is based on the marine regions and subregions under the Marine Strategy Framework Directive. Additionally, Norwegian (Barent Sea and Norwegian Sea) and Icelandic waters (’Iceland Sea’) have been added (see Surrounding seas of Europe). Note that within the North East Atlantic region only the subregions within EEZ boundaries (~200 nm) have been included. This dataset underpins the findings and cartographic representations published in the report "Contaminants in Europe's Seas" (EEA, 2019).
-

Frise chronologique des millésimes de modes d'occupation des sols produits par les régions françaises et par le programme Corine Land Cover.
-

Successive infections with Vibrio harveyi were conducted in two populations of the European abalone in order to examine which genes may be involved in improved survival to the disease in the St. Malo population.
-
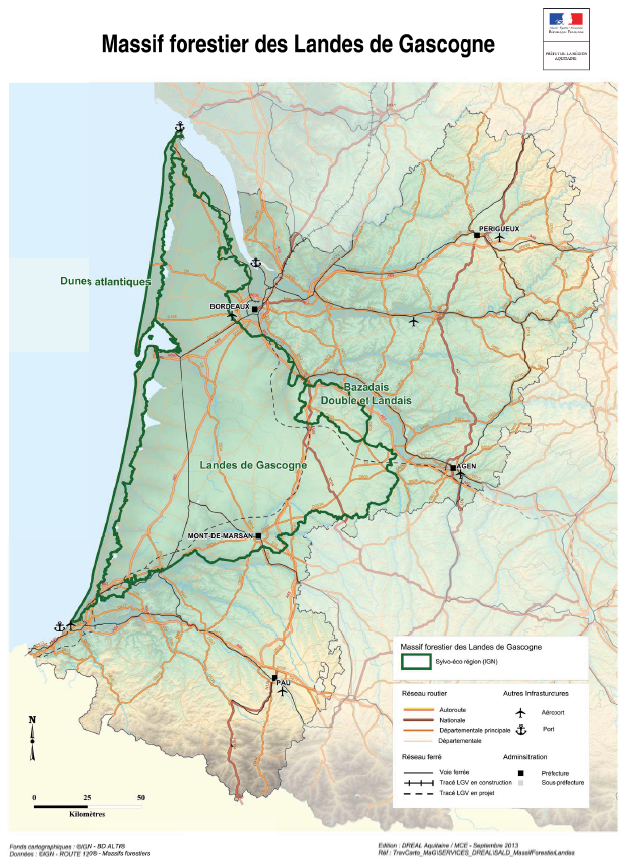
Ces travaux ont été réalisés dans le cadre de la Directive Territoriale d'Aménagement et de Développement Durable (DTADD) portée par la Préfecture de la région ex-Aquitaine. La partie I de ces travaux porte sur les valeurs du massif forestier des Landes de Gascogne. Le massif est dépositaire d’importantes valeurs et fonctions non marchandes d’intérêt général notamment : paysagères, naturalistes, hydrologiques et climatiques. Ce rapport explique également que les modes de valorisation du territoire, autres que ceux liés à la production de bois d’œuvre et d’industrie, interfèrent étroitement avec la présence même de la forêt de production : l'activité touristique, l'arrivée de nouveaux habitants et l'économie induite, ainsi que le foncier forestier.
-
List of fish stocks referenced for the year 2018. The repository includes 477 stocks. Each stock is identified by a unique key in accordance with the ICES codification in use. Each record contains a stock identifier, a species or group of species identifier according to the ASFIS/FAO classification, the English stock name, the Latin name of the species, the assessment area according to the FAO codification of fishing sectors. When the stock assessment area groups a series of sectors, the first and last sectors in the series are separated by a dash.
-
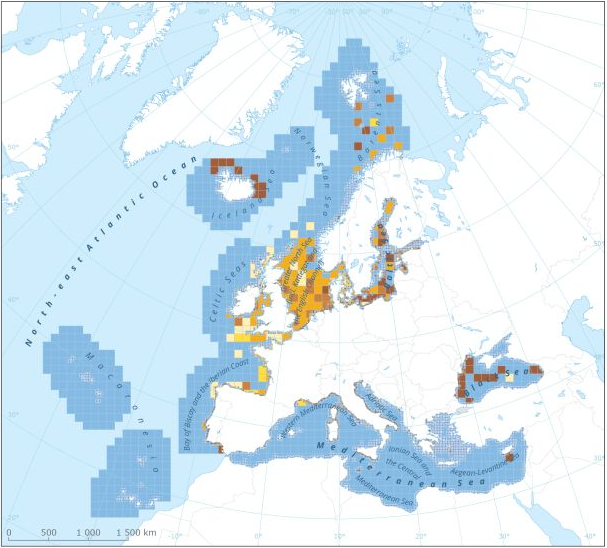
This dataset presents the resulting assessment grid (based on the EEA reference grid) with the classification of chemical status of the transitional, coastal and marine waters in the context of the Water Framework Directive (WFD) and the Marine Strategy Framework Directive (MSFD). This classification has been performed using the CHASE+ tool, with classifications of the matrices ‘water’, ‘sediment’ and ‘biota’ and indicators of ‘biological effects’, as well as an integrated classification of chemical status, combining results of all matrices. The chemical status is evaluated in five classes, where NPAhigh and NPAgood are recognised as ‘non-problem areas’ and PAmoderate, PApoor and PAbad are recognised as ‘problem areas’. This is the assessment made excluding concentrations of polybrominated diphenyl ethers (PBDEs) The overall area of interest used is based on the marine regions and subregions under the Marine Strategy Framework Directive. Additionally, Norwegian (Barent Sea and Norwegian Sea) and Icelandic waters (’Iceland Sea’) have been added (see Surrounding seas of Europe). Note that within the North East Atlantic region only the subregions within EEZ boundaries (~200 nm) have been included. This dataset underpins the findings and cartographic representations published in the report "Contaminants in Europe's Seas" (EEA, 2019): https://www.eea.europa.eu/publications/contaminants-in-europes-seas.
 Catalogue PIGMA
Catalogue PIGMA