2022
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Service types
Scale
Resolution
-

This dataset consists of metatranscriptomic sequencing reads corresponding to coastal micro-eukaryote communities sampled in Western Europe in 2018 and 2019.
-
Serveur wms du projet CHARM II
-
Serveur wms du projet CHARM III
-
In recent years, large datasets of in situ marine carbonate system parameters (partial pressure of CO2 (pCO2), total alkalinity, dissolved inorganic carbon and pH) have been collated. These carbonate system datasets have highly variable data density in both space and time, especially in the case of pCO2, which is routinely measured at high frequency using underway measuring systems. This variation in data density can create biases when the data are used, for example for algorithm assessment, favouring datasets or regions with high data density. A common way to overcome data density issues is to bin the data into cells of equal latitude and longitude extent. This leads to bins with spatial areas that are latitude and projection dependent (eg become smaller and more elongated as the poles are approached). Additionally, as bin boundaries are defined without reference to the spatial distribution of the data or to geographical features, data clusters may be divided sub-optimally (eg a bin covering a region with a strong gradient). To overcome these problems and to provide a tool for matching in situ data with satellite, model and climatological data, which often have very different spatiotemporal scales both from the in situ data and from each other, a methodology has been created to group in situ data into ‘regions of interest’, spatiotemporal cylinders consisting of circles on the Earth’s surface extending over a period of time. These regions of interest are optimally adjusted to contain as many in situ measurements as possible. All in situ measurements of the same parameter contained in a region of interest are collated, including estimated uncertainties and regional summary statistics. The same grouping is done for each of the other datasets, producing a dataset of matchups. About 35 million in situ datapoints were then matched with data from five satellite sources and five model and re-analysis datasets to produce a global matchup dataset of carbonate system data, consisting of 287,000 regions of interest spanning 54 years from 1957 to 2020. Each region of interest is 100 km in diameter and 10 days in duration. An example application, the reparameterisation of a global total alkalinity algorithm, is shown. This matchup dataset can be updated as and when in situ and other datasets are updated, and similar datasets at finer spatiotemporal scale can be constructed, for example to enable regional studies. This dataset was funded by ESA Satellite Oceanographic Datasets for Acidification (OceanSODA) project which aims at developing the use of satellite Earth Observation for studying and monitoring marine carbonate chemistry.
-

Phenotypic plasticity, the ability of a single genotype to produce multiple phenotypes, is important for survival when species are faced with novel conditions. Theory predicts that range-edge populations will have greater phenotypic plasticity than core populations, but empirical examples from the wild are rare. The honeycomb worm, Sabellaria alveolata (L.), constructs the largest biogenic reefs in Europe, which support high biodiversity and numerous ecological functions. In order to assess the presence, causes and consequences of intraspecific variation in developmental plasticity and thermal adaptation in the honeycomb worm, we carried out common-garden experiments using the larvae of individuals sampled from along a latitudinal gradient covering the entire range of the species. We exposed larvae to three temperature treatments and measured phenotypic traits throughout development. We found phenotypic plasticity in larval growth rate but local adaptation in terms of larval period. The northern and southern range-edge populations of S. alveolata showed phenotypic plasticity for growth rate: growth rate increased as temperature treatment increased. In contrast, the core range populations showed no evidence of phenotypic plasticity. We present a rare case of range-edge plasticity at both the northern and southern range limit of species, likely caused by evolution of phenotypic plasticity during range expansion and its maintenance in highly heterogeneous environments. This dataset presents the raw image data collected for larval stages of Sabellaria alveolata from 5 populations across Europe and Northern Africa, exposed to 15, 20 and 25 C. Included are also opercular crown measurements used to estimate de size classes of individuals present in each population. All measurements made with the images collected are presented in an Excel spreadsheet, also available here.
-
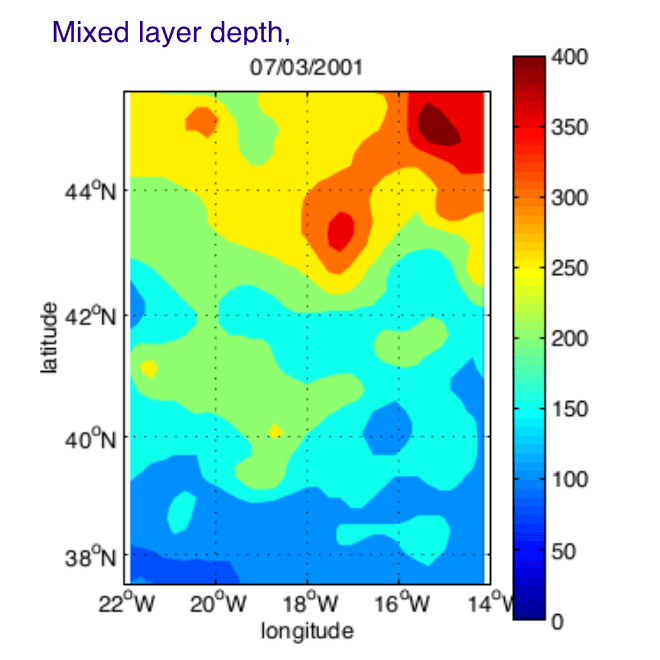
The Programme Ocean Multidisciplinaire Meso Echelle (POMME) was designed to describe and quantify the role of mesoscale processes in the subduction of mode waters in the Northeast Atlantic. Intensive situ measurements were maintained during 1 year (September 2000 - October 2001), over a 8 degrees square area centered on 18 degrees W, 42 degrees N. In order to synthesized the in-situ physical observations, and merge them with satellite altimetry and surface fluxes datasets, a simplified Kalman filter has been designed. Daily fields of temperature, salinity, and stream function were produced on a regular grid over a full seasonal cycle. We propose here the gridded fields (KA_ files) and the in-situ datasets used by the analysis (Data_ files).
-
Serveur wms sur les photos anciennes
-

Understanding the dynamics of species interactions for food (prey-predator, competition for resources) and the functioning of trophic networks (dependence on trophic pathways, food chain flows, etc.) has become a thriving ecological research field in recent decades. This empirical knowledge is then used to develop population and ecosystem modelling approaches to support ecosystem-based management. The TrophicCS data set offers spatialized trophic information on a large spatial scale (the entire Celtic Sea continental shelf and upper slope) for a wide range of species. It combines ingested prey (gut content analysis) and a more integrated indicator of food sources (stable isotope analysis). A total of 1337 samples of large epifaunal invertebrates (bivalve mollusks and decapod crustaceans), zooplankton, fish and cephalopods, corresponding to 114 species, were collected and analyzed for stable isotope analysis of their carbon and nitrogen content. Sample size varied between taxa (from 1 to 52), with an average of 11.72 individuals sampled per species, and water depths ranged from 57 to 516 m. The gut contents of 1026 fish belonging to ten commercially important species: black anglerfish (Lophius budegassa), white anglerfish (Lophius piscatorius), blue whiting (Micromesistius poutassou), cod (Gadus morhua), haddock (Melanogrammus aeglefinus), hake (Merluccius merluccius), megrim (Lepidorhombus whiffiagonis), plaice (Pleuronectes platessa), sole (Solea solea) and whiting (Merlangius merlangus) were analyzed. The stomach content data set contains the occurrence of prey in stomach, identified to the lowest taxonomic level possible. To consider potential ontogenetic diet changes, a large size range was sampled. The TrophicCS data set was used to improve understanding of trophic relationships and ecosystem functioning in the Celtic Sea. When you use the data in your publication, we request that you cite this data paper. If you use the present data set (TrophicCS) for the majority of the data analyzed in your study, you may wish to consider inviting at least one author of the core team of this data paper to become a collaborator /coauthor of your paper.
-

Particularly suited to the purpose of measuring the sensitivity of benthic communities to trawling, a trawl disturbance indicator (de Juan and Demestre, 2012, de Juan et al. 2009) was proposed based on benthic species biological traits to evaluate the sensibility of mega- and epifaunal community to fishing pressure known to have a physical impact on the seafloor (such as dredging and bottom trawling). The selected biological traits were chosen as they determine vulnerability to trawling: mobility, fragility, position on substrata, average size and feeding mode that can easily be related to the fragility, recoverability and vulnerability ecological concepts. The five categories retained are functional traits that were selected based on the knowledge of the response of benthic taxa to trawling disturbance (de Juan et al., 2009). They reflect respectively the possibility to avoid direct gear impact, to benefit from trawling for feeding, to escape gear, to get caught by the net and to resist trawling/dredging action, each of these characteristics being either advantageous or sensitive to trawling. To expand this approach to that proposed by Certain et al. (2015), the protection status of certain species was also indicated. To enable quantitative analysis, a score was assigned to each category: from low sensitivity (0) to high sensitivity (3). Biological traits of species have been defined, from the BIOTIC database (MARLIN, 2014) and from information given by Garcia (2010), Le Pape et al. (2007) and Brind’Amour et al. (2009). For missing traits, additional information from literature has been considered. The protection status of each taxa was also scored: Atlantic species listed in OSPAR List of Threatened and/or Declining Species and Habitats (https://www.ospar.org/work-areas/bdc/species-habitats/list-of-threatened-declining-species-habitats) and Mediterranean species listed in Vulnerable Marine Ecosystems (FAO, 2018 and Oceana, 2017) were scored 3 and other species were scored 1. The scores of 1085 taxa commonly found in bottom trawl by-catch in the southern North Sea, English Channel and north-western Mediterranean were described.
-
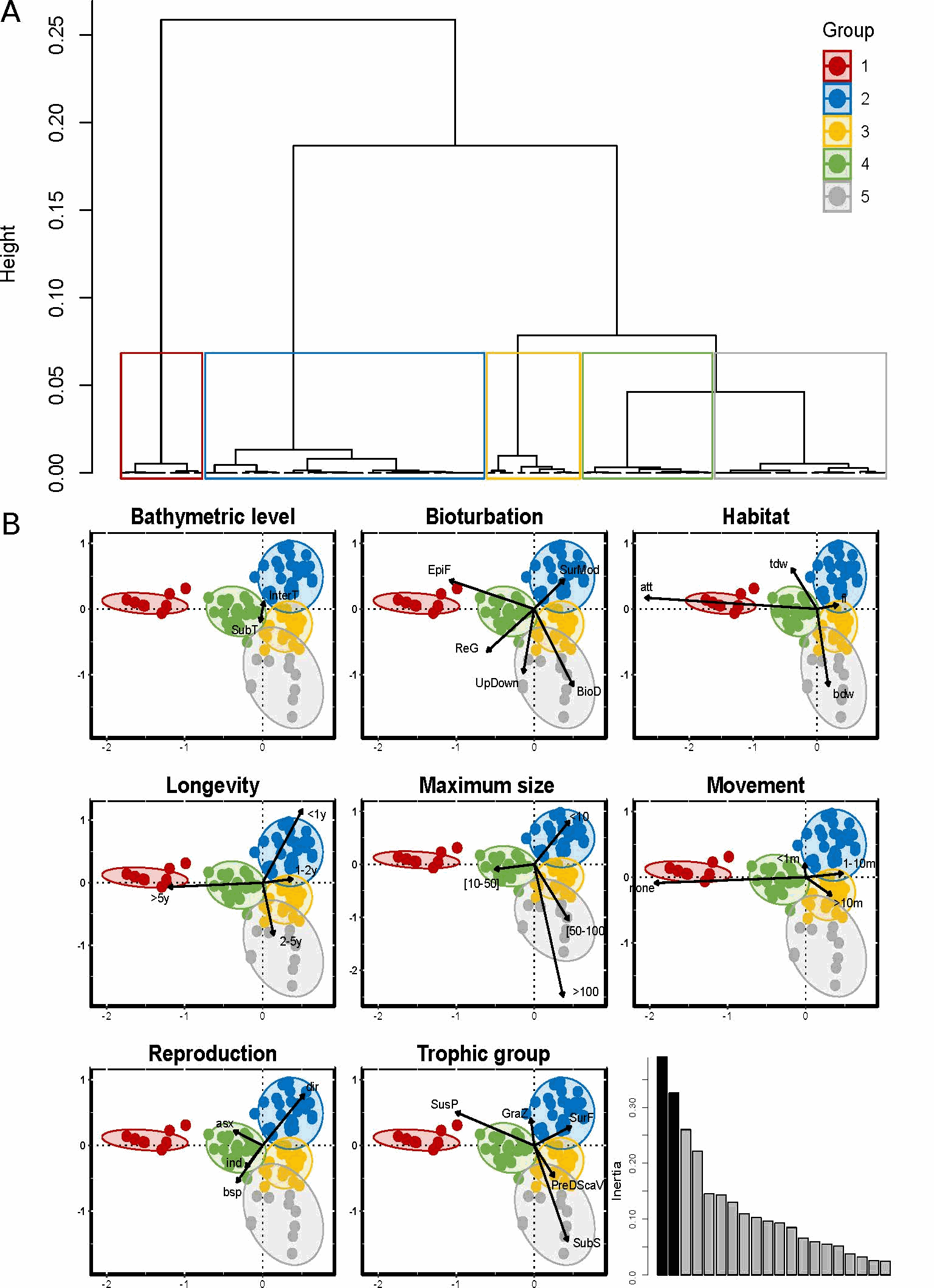
Reef-building species are recognized as having an important ecological role and as generally enhancing the diversity of benthic organisms in marine habitats. However, although these ecosystem engineers have a facilitating role for some species, they may exclude or compete with others. The honeycomb worm Sabellaria alveolata (Linnaeus, 1767) is an important foundation species, commonly found from northwest Ireland to northern Mauritania (Curd et al., 2020), whose reef structures increase the physical complexity of the marine benthos, supporting high levels of biodiversity. Local patterns and regional differences in taxonomic and functional diversity were examined in honeycomb worm reefs from ten sites along the northeastern Atlantic to explore variation in diversity across biogeographic regions and the potential effects of environmental drivers. To characterize the functional diversity at each site, a biological trait analysis (BTA) was conducted (Statzner et al., 1994). Here we present the functional trait database used for the benthic macrofauna found to live in association with honeycomb worm reefs. Eight biological traits (divided into 32 modalities) were selected (Table 1), providing information linked to the ecological functions performed by the associated macrofauna. The selected traits provide information on: (i) resource use and availability (by the trophic group of species, e.g. Thrush et al. 2006); (ii) secondary production and the amount of energy and organic matter (OM) produced based on the life cycle of the organisms (including longevity, maximum size and mode of reproduction, e.g. (Cusson and Bourget, 2005; Thrush et al., 2006) and; (iii) the behavior of the species in general [i.e. how these species occupy the environment and contribute to biogeochemical fluxes through habitat, movement, and bioturbation activity at different bathymetric levels, e.g. (Solan et al., 2004; Thrush et al., 2006; Queirós et al., 2013). Species were scored for each trait modality based on their affinity using a fuzzy coding approach (Chevenet et al., 1994), where multiple modalities can be attributed to a species if appropriate, and allowed for the incorporation of intraspecific variability in trait expression. The information concerning polychaetes was derived primarily from Fauchald et al (1979) and Jumars et al (2015). Information on other taxonomic groups was obtained either from databases of biological traits (www.marlin.ac.uk/biotic) or publications (Naylor, 1972; King, 1974; Caine, 1977; Lincoln, 1979; Holdich and Jones, 1983; Smaldon et al., 1993; Ingle, 1996; San Martín, 2003; Southward, 2008; Gil, 2011; Leblanc et al., 2011; Rumbold et al., 2012; San Martín and Worsfold, 2015; Jones et al., 2018). Map indicating the locations of the 10 study sites in the UK, France and Portugal within the four biogeographic provinces defined by Dinter (2001). (All sites were sampled in 8 different stations, except for UK4 where 5 stations were sampled).
 Catalogue PIGMA
Catalogue PIGMA